Resources
We are actively engaged in generating tools for the identification and analysis of neuronal circuits in the fly. We have developed techniques for the constitutive suppression (White et. al., 2001 ) and enhancement of neuronal excitability (Luan, Lemon et al., 2006 ; Nitabach et al. 2006 ), as well as a tool for acutely activating neurons in response to small temperature shifts (Peabody et al., 2009 ).
We have also developed general methods for targeting genetic manipulations to cell types of interest. These include the intersectional Split Gal4 system (Luan et al., 2006 ; Luan et al.,2020 ), which permits refined targeting of transgenes to small subsets of neurons and has been augmented by our recently introduced suppressor of Split Gal4 activity, the Killer Zipper (Dolan, Luan et al., 2017 ). In addition, we have developed the SpaRCLIn method, which permits stepwise refinement of genetic targeting to cell types of interest using a split cre recombinase (Luan et al.,2020 ).
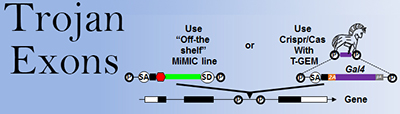
To target neurons that express a given gene of interest, we have taken advantage of the ribosomal skipping mechanism of viral 2A peptides to create multiple tools. The first of these was the T2A-Gal4 In-Frame-Fusion method (“T2A-GIFF;” Diao and White, 2012 ), which was followed by the more versatile and modular technology of “Trojan exons” (Diao et al., 2015 ). Trojan exons are interchangeable synthetic exons that are preceded by a universal splice acceptor and which can be inserted in-frame into a coding intron in any gene of interest. Cells expressing a gene can thus be targeted with high fidelity.
All of our published plasmids and fly lines are freely available, most of them from public repositories. Our published constructs for the Split Gal4 system, Trojan exons, Killer Zipper, and various effectors, such as EKO and NaChBac are available through both Addgene and the Drosophila Genomic Resources Center . If you are unable to find what you want, please contact us. Many of our published fly lines (particularly those that represent commonly used genetic tools) have been deposited at the Bloomington Drosophila Stock Center where they are publicly available. Other lines created by the lab are available upon request.

